Research

EK-Br3 crystals grown by Kailey Martin.
Exploration of 3D conformational space occupied by quadruplex DNA and i-motifs
G-quadruplexes and i-motif structures are promising drug targets for cancer. Due to their high structural diversity and a limited number of available high-resolution structures, our ability to design potent, selective ligands is hampered. Structural information on the i-motif is extremely scarce. To develop structure-selective ligands as drug candidates for cancer intervention, high-resolution structural information is necessary. Driven by this gap in knowledge we are working toward crystallization of : 1) a variety of oncogene promoter G-quadruplexes in complex with ligands; 2) i-motif DNA from oncogene promoters alone and in complex with ligands; 3) repetitive DNA associated in replication stress; 4) DNA repeat from centromeres 2 and 3; 5) Mitochondrial G-quadruplex DNA.
Interaction of G-quadruplex DNA with small molecule ligands
Our laboratory demonstrated that N-methylmesoporphyrin IX (NMM) has excellent selectivity for G-quadruplex vs dsDNA. Our interest in NMM was sparked by reports of its biological utility and widespread applications. We have characterized the interactions of NMM with a variety of G-quadruplex DNA folds and have determined its crystal structures in complex with human telomeric DNA and Tetrahymena thermophila telomeric DNA. NMM provides an excellent platform for investigating the structural and chemical requirements for highly selective G-quadruplex binders. Currently we are working on a number of ligands from Anton Granzhan's lab from Université Paris Cité. Those ligands include PyDH2, TrisQ, PhenDH2 and PhenDH9. Other well-studied and highly selective ligands include PDS, Braco-19, 360A, RHPS4, and PhenDC3. Structural characterization of their binding to G-quadruplexes will inform the design of improved analogues.
Funding
- NIH 1R15 (AREA) Deciphering the structure and dynamics of non-canonical DNA implicated in cancer; 2020-2023. No cost extension to 2024
- (co-PI) NIH R01 Mitochondrial G-quadruplex structures in health and disease, 2019-2024 (PI: Brett Kaufman)
Collaborators
We actively collaborate with US and international researchers whose expertise and resources complement our work and provide extraordinary opportunities for the students. Our current collaborators include J.-L. Mergny (Ecole Polytechnique), Anton Granzhan (Université Paris Cité), Liliane Mouawad (Institute Curie), Brahim Heddi (Université Paris-Saclay ), Antonio Randazzo (University of Naples Federico II) and Stephen Neidle (University College London). Liliya spent the 2010-11 and 2014-15 academic years on sabbatical in the laboratory of J.-L. Mergny at the European Institute of Chemistry and Biology (IECB) in France, and 2018-2019 sabbatical in the laboratory of J. Plavec (National Slovenian NMR center) and V. Gabelica (IECB). Through these collaborations, five of my students and I learned new biophysical, biological, and structural techniques for characterizing non-canonical DNA structures. I, along with one of my students, also spent a few weeks in the laboratory of B. Chaires learning calorimetric techniques.

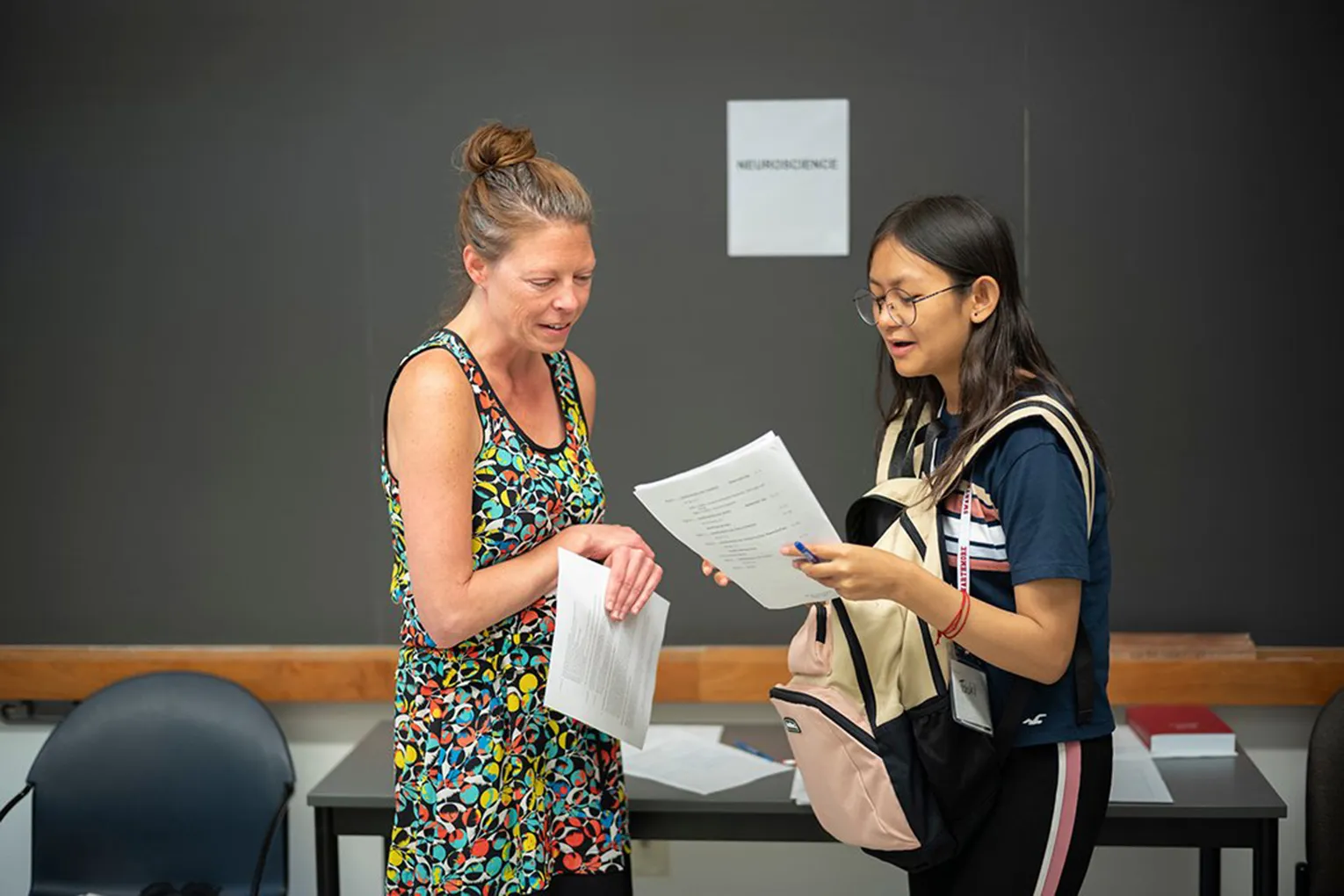
from New York University School of Medicine, visiting Swarthmore to learn
about our lab's biophysical characterization techniques

Dr. Brad Chaires (University of Louisville), and Barrett Powell '18 at
the International Meeting on Quadruplex Nucleic Acids in Prague, June 2017

Joint group meeting with Dr. Eric Brown (far left) from the University of Pennsylvania

Thao Tran (IECB, Bordeaux), F. Brad Johnson (UPenn Med School)
Jack Nicoludis '12 and Liliya Yatsunyk at the Third International Meeting on
G-quadruplex DNA and G-assembly in Italy, Sorrento, July 2011
Undergraduate research
Most research is our laboratory is done by Swarthmore undergraduates. Students with interest in cancer research, crystallography, synthetic work, biophysical work, interdisciplinary work between chemistry, biology, and physics are highly encouraged to apply. Students from the departments other than Chemistry are also encouraged to apply. Our lab is a home for Engineering, Biology, and Psycology students. Based on background and interests, students will pursue independent projects or be teamed with senior students in the lab. Contact Professor Yatsunyk via e-mail or stop by our lab and talk to students about their experience.